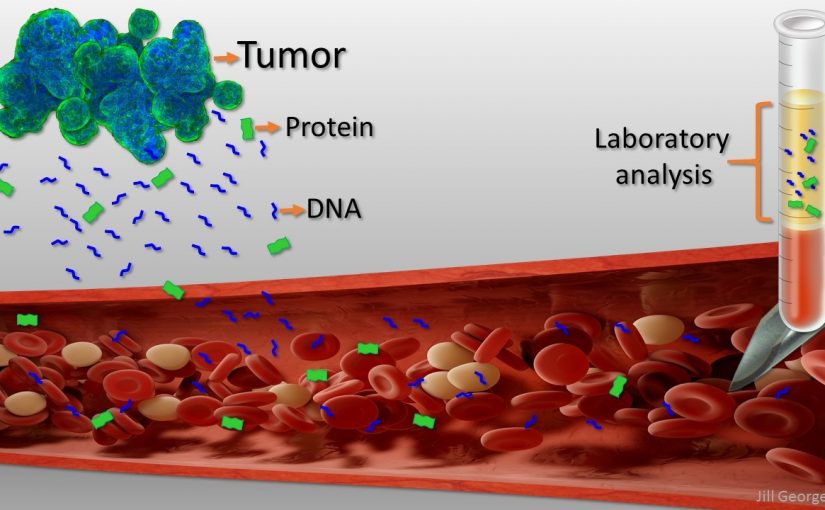
DNA shed by tumors into the bloodstream is starting to be used for the early detection of cancer and for monitoring how well treatment is working. Such DNA, which is called circulating tumor DNA (ctDNA), hasn’t been quite as helpful with respect to another important task—the selection of the best available anticancer treatment. In fact, the use of ctDNA in guiding treatment decisions in solid cancers has been controversial.
Optimism that ctDNA can guide treatment decisions may be justified, suggest scientists based at the University of Helsinki. These scientists, led by Sampsa Hautaniemi, PhD, professor of systems biology, reported that they were able to use ctDNA shed by ovarian tumors to identify clinically actionable mutations.
Hautaniemi’s research group has implemented a clinical circulating tumor DNA workflow to detect clinically actionable alterations in more than 500 cancer-related genes. In addition, the group analyzed 78 circulating tumor DNA samples from 12 ovarian cancer patients before, during, and after the treatment. The data the researchers collected were analyzed using an open-access ctDNA-tailored bioinformatics analysis pipeline and in-house Translational Oncology Knowledgebase to detect clinically actionable genomic alterations.
The results of this work appeared May 3 in JCO Precision Oncology, in an article titled, “Prospective Longitudinal ctDNA Workflow Reveals Clinically Actionable Alterations in Ovarian Cancer.”
“Our results show good concordance of mutations and copy number alterations in ctDNA and tumor samples, and alterations associated with clinically available drugs were detected in seven patients (58%),” the article’s authors indicated. “Treatment of one chemoresistant patient was changed on the basis of detection of ERBB2 amplification, and this ctDNA-guided decision was followed by significant tumor shrinkage and complete normalization of the cancer antigen 125 tumor marker.”
The current study focused on high-grade serous ovarian cancer (HGSOC), the most common and aggressive form of epithelial ovarian cancer. Over half of the patients diagnosed with HGSOC die within five years of diagnosis, that is, annually more than 150,000 women globally.
HGSOC tumors consist of several heterogeneous cell populations with a large number of mutations. This genetic variability makes it difficult to find drugs that would kill all the cancer cells, and to which the cells would not become resistant during treatment. To improve the efficacy of HGSOC treatment, the current study evaluated whether ctDNA, as a precision medicine modality, could be used to select the best available treatment.
“Analyzing ctDNA enables us to detect genomic alterations [not only in early-stage cancers, but] also in late-stage cancers, [when] taking biopsies from the tumor is difficult or even impossible,” said Hautaniemi. “In the current research, we demonstrated that circulating tumor DNA can be used to monitor the patient responses to treatment, and to find drug combinations to target the Achilles’ heels of the tumor’s genomic profile.”
Hautaniemi’s research group has implemented a clinical circulating tumor DNA workflow to detect clinically actionable alterations in more than 500 cancer-related genes. In the newly published article, the authors assert that its proof-of-concept study demonstrates that ctDNA can guide clinical decisions. Furthermore, the researchers say their results show that longitudinal ctDNA samples can be used to identify poor-responding patients after first cycles of chemotherapy. Finally, the researchers maintain that they have provided the first comprehensive, open-source circulating tumor DNA workflow for detecting clinically actionable alterations in solid cancers.
“The prognosis of ovarian cancer patients is still poor,” Hautaniemi pointed out. “However, the future looks brighter as research has discovered targeted therapies for the genomic alterations also in ovarian cancer tumors. We are currently in the forefront of the precision medicine for ovarian cancer. Our goal is to find out how ovarian cancer becomes resistant to current treatments, and to use this knowledge in finding more effective precision treatments.”
This article was published by Genetic Engineering & Biotechnology News.